
Data Analysis Fusion Team
The Fourth Methodology (Data Analysis Fusion): Transforming
Biology into a Predictable Science

Data Analysis Fusion Team
Team Leader
Satoru MIYANO
I may face scorn for saying this, but biology really is the only field of science that does not have a "scientific" language. Practically speaking, no discoveries of what can be called a principle have been made and researchers continue to basically present facts similar to theorems in Mathematics. For example, we may prove with a molecular biology method that a certain gene controls a particular gene cluster and, as a result, can be identified as the cause of a particular phenotype. We write about facts that support such findings. However, while this may be fascinating in its own right, we are basically just letting the world know we have discovered this or that. With this approach, even after a thousand years, life science would not evolve to be a "predictable science."
In 2003, the National Institute of Health (NIH) in the U.S. launched a roadmap for biomedical research to fully benefit from the completion of the human genome project. It includes the following sentence: "All of these techniques generate large amounts of data and biology is fast changing into a science of information management." At the time of publication, biologists and medical researchers were bewildered by the message. Now, however, everyone strongly recognizes this to be a reality. That is, although data generation continues to be important, the focus of biology as a science is shifting toward how to analyze and interpret vast amounts of data. Concurrent with efforts to effectively deal with huge quantities of facts, there is also a need to change biology into a "predictable science." An acknowledgement of the importance of computational science in the life sciences reflects this need. Therefore, it is important to pursue the development of life science software that uses next-generation supercomputers, which will lead to the establishment of an information infrastructure to make this a reality.
Our Data Analysis Fusion Team is divided into four groups, each with its own theme: "estimations and applications of large-scale genetic networks," "development of new algorithms that relate to large-scale genome polymorphism data and phenotypic data and a review of the validity and usability of these algorithms," "estimations and applications of large-scale protein networks" and "development of data assimilation technologies for the simulation of living matter." The groups work synergistically to establish more sophisticated modeling technologies by analyzing networks of molecular interactions based on gene expression data, etc., then combining the extracted models with estimated dynamics.
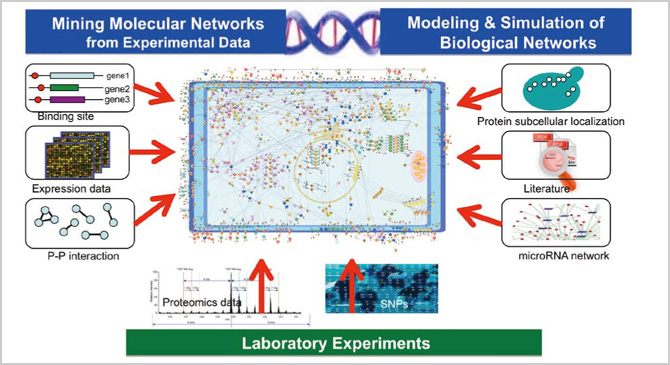
Here is a simple overview on our research and development. We first predict a large-scale molecular network by computing data such as gene expression data. Using the results as a map, we then look for molecular interaction or genes that may be relevant to drugs and diseases. In the field of geophysics, for example, exploration by artificial satellites made global earth maps readily available and allowed researchers to keep track of temperature and humidity data. Likewise, in life science, DNA chips allowed us to identify the gene expression of a particular cell at a glance. With such information, we can ascertain the correlation between certain molecules. In other words, we are developing a map of a dynamic network, which is to be further refined by identifying where the gene responsible for a certain disease and its molecular interaction are located on the map, in order to develop an even more sophisticated model. Due to the number of variations of phenomena in an actual human body, however, a standard model is not always applicable. We could overcome such issues by employing data assimilation techniques. This approach will allow us to create a personalized map in which the individual's information is incorporated into a standard model based on molecular-biological facts along with networks and dynamics from a vast amount of data.
What's important here is that as long as a biologically-sound standard model exists, it is possible to then build a specific model for an individual by applying personal data, even with the existence of many variables. Building a tailor-made, personal model from scratch could take an untold number of years. If we can make such a model with the use of data assimilation, it will allow us to conduct simulations to identify which drugs are needed for each patient and open up new possibilities in therapy.
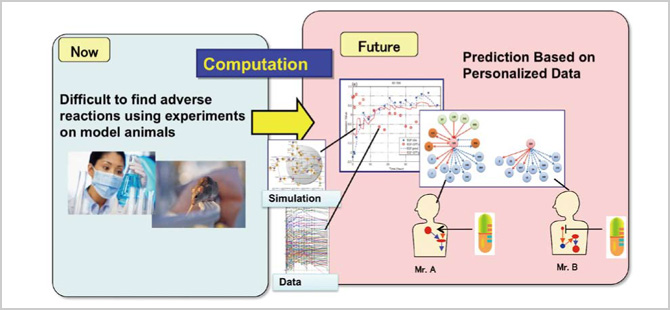
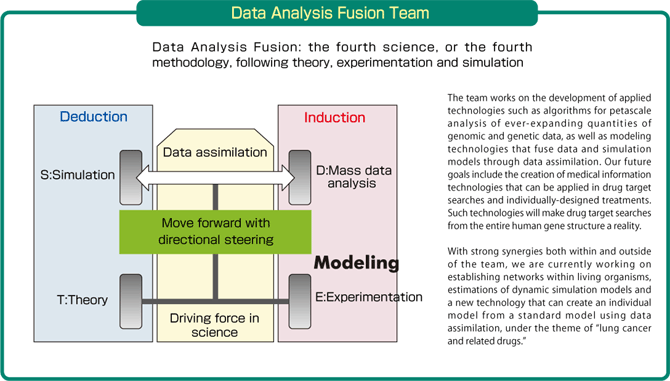
Our new approach, "data analysis fusion," integrates deduction (i.e., simulations) and induction (i.e., mass data processing) through data assimilation. It is the fourth science, or the fourth methodology, a new addition to the three pillars of science: theory, experimentation and simulation. In addition, under the theme of "lung cancer and related drugs," we are currently developing a new technology that can create an individual model from a standard model using the state-space modeling method
BioSupercomputing Newsletter Vol.1
- SPECIAL INTERVIEW
- Innovative Approach for Understanding Phenomena of Life Exploring New Possibilities with Bio-supercomputing
Computational Science Research Program Deputy Program Director Ryutaro HIMENO
- A Message from the Team Leader
- Simulations to Understand the Functions of the Biopolymers that Play Fundamental Roles in Life
Molecular Scale Team Team Leader Akinori KIDERA - Develop a 3-D Model of the Entire Human Body and Understand In Vivo Phenomena to Utilize for Medical Purposes
Organ and Body Scale Team Team Leader Shu TAKAGI - The Fourth Methodology (Data Analysis Fusion): Transforming Biology into a Predictable Science
Data Analysis Fusion Team Team Leader Satoru MIYANO
- Report on Research
- Prediction of Transmembrane Dimer Structure of Amyloid Precursor Protein using Replica-Exchange Molecular Dynamics Simulations
Molecular Scale Team Naoyuki MIYASHITA / RIKEN Advanced Science Institute (Molecular Scale WG) Yuji SUGITA - Simulation for Charged Particle Therapy
Organ and Body Scale Team Kenichi L. ISHIKAWA - Prospects of Prognostic Prediction Based on Genome-wide Association Study and Genetic/Non-genetic Factors
Riken Center for Genomic Medicine (Data Analysis Fusion WG) Naoyuki KAMATANI - Key Technology Supporting Petascale Computing
High-performance Computing Team Kenji ONO / Satoshi ITO / Daisuke WATANABE
