 Strategic Programs for Innovative Research
Strategic Programs for Innovative Research
Field 1 Supercomputational Life Science
Theme 3 Hierarchical integrated simulation for predictive medicine
Aiming to realize hierarchical integrated
simulation in the circulatory organ system and
the musculoskeletal / cerebral nervous systems

Professor, Department of Mechanical Engineering and Department of
Bioengineering The University of Tokyo
Shu Takagi
(Theme3 GL)
●Elucidate life systems in top-down approach
The ultimate goal of Theme 3 is to elucidate the meaning of “being alive.” To this end, we will work on hierarchical integrated simulation, which forms a hierarchical human body model, with the aim to understand life as an integrated system.
Our lives are sustained by ingesting nutrition from the mouth, which is transported throughout the body via the blood and by breathing air to obtain oxygen in the lungs, which is transported throughout the human body again via the blood. Through these processes, the cells constituting the human body play their respective functions with the nutrition and oxygen delivered to them. After this process, a direction for further nutrient intake will be transmitted to our brain and our brain will feel hungry and encourage us to eat more. There are naturally various smaller processes, which are required for the sustainability of life, with a system that transfers heat, mass, and information in various forms. In plain words, Theme 3 aims to elucidate life systems in such an approach as explaining what is happening in the body with the above phenomena as a starting point.
For example, when a disease was cured by an effective medicine, it is correct to explain, on the molecular level, that “the disease was cured as a result of normal cellular functioning with a certain function of protein inhibited by the medicine.” If the disease was not cured, however, there are many cases where the reason cannot be explained, only locally on the molecular level, why the disease was not cured. In fact, there are many medicines that are not effective although they should take effect on the molecular level. When symptoms of a disease disappear, your brain will feel that “the disease has been cured.” In that case, “the cure of your disease” is felt in the brain finally via the nervous system after your cells normally function with the protein functions changed, and then your tissues and organs are normally activated as populations of such cells rather than the individual functioning of proteins or cells. In other words, outcomes are mostly not determined only by local events. Accordingly, if a medicine is not effective, you should think of other various factors involved. In order to understand “the meaning of being alive,” various factors need to be grasped from the molecular level to the systemic level. It is difficult, however, to elaborately simulate all factors from the molecular level to the systemic level and there are inevitable limitations. What is important is the methodology as to how the hierarchies should be integrated.
There are two approaches to the understanding of life phenomena where various events are inter-related in a complicated manner both in terms of scale and time. One is a bottom-up approach that builds up small factors
to lead to the understanding of larger phenomena, and the other is a top-down approach that starts with a condition, where the whole body is well-maintained, and goes down to smaller factors to explore how the small factors function and what is effective in maintaining the whole body. The Theme 3 Team aims to elucidate life phenomena with the latter approach.
Our body is equipped with a function called homeostasis that tries to restore to a normal condition. Thanks to homeostasis, a slightly disturbed condition will not aggravate immediately. The condition will drastically become pathological, however, when it exceeds a certain level. Although it may be difficult to grasp quantitatively what significant changes will occur at which stages due to what effective functions, we believe that a top-down approach is better to find what significant changes will occur at what levels of which factors. On the contrary, it is very difficult in a bottom-up approach to find what factor remains last. Hierarchical integration is coarse graining in the sense that it tries to focus on different hierarchies by cutting off something. It should be noted that simple averaging will erase the functions and features that may appear in the next hierarchies. What should be left depends on the functions and features that will appear in upper hierarchies in the end. Which factors should be linked to what parts and how they can be linked together to leave cardiac functions for the heart and leave blood flow functions for blood flow properly based on the theories of the lower hierarchies ─ it is no exaggeration to say that most of the study subjects, which we have dealt with so far, focused on the handling of this point
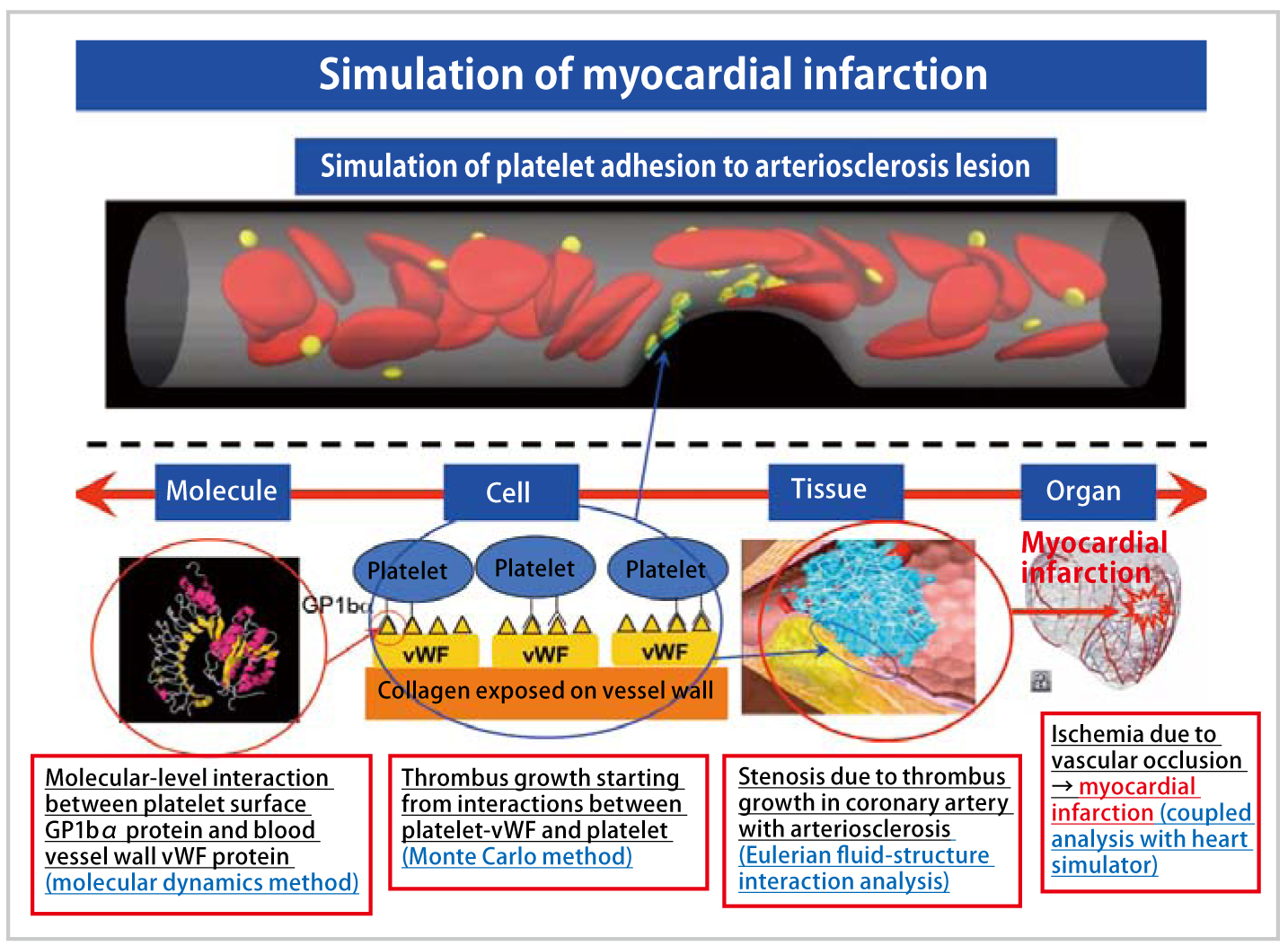
We will aim to reproduce the scenario leading to myocardial infarction by integrating the thrombosis simulator and the heart simulator. We will reproduce vascular occlusion in a contracting/dilating coronary artery through the coupled analysis of heartbeat by further expanding the multi-scale thrombus simulator, incorporating the previous simulation ranging from protein-molecular-level interactions to fluid-dynamicslevel blood flow phenomena.
●Exploit the results of the Grand Challenge Program for further development
In the Grand Challenge Program, which has been carried out for the past five years, various living body simulators with excellent performance have been developed such as the thrombus simulator, the heart simulator, the musculoskeletal simulator, and the cerebral nervous system simulator. In Theme 3, we wish to improve the infrastructure tool for reproducing complicated processes relating to various diseases by integrating these simulators, which have been developed with the “K computer” on an individual basis so far, by positively exploiting the previous results with an eye to developing the efforts into construction of frames for integration into a hierarchical human whole-body simulation in future. In addition, we will work on applying disease-related simulation results to the prediction of pathological conditions, and support for treatment.
We selected myocardial infarction and Parkinson disease as two specific simulation targets. As mentioned above, the circulatory organ system is a mass transfer network and the nervous system is a signal transduction network, which are vital in the human body for transfer of heat, mass, and information for the sustainability of life, and function as what may be called human body trunk line networks. The circulatory organ system and the nervous system are also very important for future construction of a systemic integrated simulator. With such importance taken into consideration, we selected myocardial infarction and Parkinson disease as targets.
In the simulation of myocardial infarction, we aim to reproduce not only a primary thrombus (platelet thrombus), where platelet adhesion and aggregation occurs in the initial stage of thrombus formation, but also the scenario, in which the growth of a secondary thrombus (fibrin thrombus) initiated by blood coagulation function develops into vascular occlusion, by expanding the thrombus simulator developed in the Grand Challenge Program. In addition, we will perform coupled analysis of the heart simulator developed by Toshiaki Hisada (The University of Tokyo) et al., to simulate the thrombus growth process from thrombotic adhesion to arteriosclerotic lesions in the coronary artery surrounding the heart to myocardial infarction with the effects of the heart simulator incorporated into the simulation result. We also wish to provide important findings useful for the development of new drugs by evaluating the effects of thrombus-related drugs.
We also aim to reproduce the simulation of Parkinson disease as an integral part of the musculoskeletal simulator and the cerebral nervous system simulator. Parkinson disease is considered to cause limb tremor and physical rigidity even in a healthy musculoskeletal body due to abnormal signals transmitted from the brain. We want to contribute to the pathological prediction and treatment support for Parkinson disease, which is one of the functional motility disorders resulting from a cranial nerve disease, by reproducing a pathology called tremor that spike signals from the brain transmit to the muscle fibers via motor neurons and cause tremors, and a pathology called rigidity that stiff muscles cause a rigid posture.
●Promote studies with an eye to the construction of human whole-body integrated simulator
In addition to the ongoing simulation of myocardial infarction and Parkinson disease, we also intend to work on the construction of infrastructure software for an integrated simulator that can be applied to a wide variety of diseases by integrating musculoskeletal and circulatory organ simulators via the nervous system. For the heart part, various diseases are expected to be simulated to study therapeutic methods and evaluate efficacy including abnormal cardiac functions due to myocardial infarction, angina pectoris, dilated cardiomyopathy, hypertrophic cardiomyopathy, and microvascular damage, and influence of stress and excitement. For Parkinson disease, we wish to develop the simulation into prediction of an optimum prescription such as concomitant use of electric stimulation with dosing based on data of individual patients on symptoms and motor tests, for example, which are different from the conventional symptomatic treatment with electric stimulation, by elucidating the mechanism and evaluating therapeutic methods through a whole-body integrated simulation including the cerebral nervous system, with consideration given to the effects of mass and signal transfer on the neuron level and the effects of signal transduction. In addition, we believe that the use of such a simulator will enable the analysis of a wide range of phenomena such as prediction of changes in body balance and damage in the event of overturning, and the difference in the effects of muscular or brain fatigue in the future.
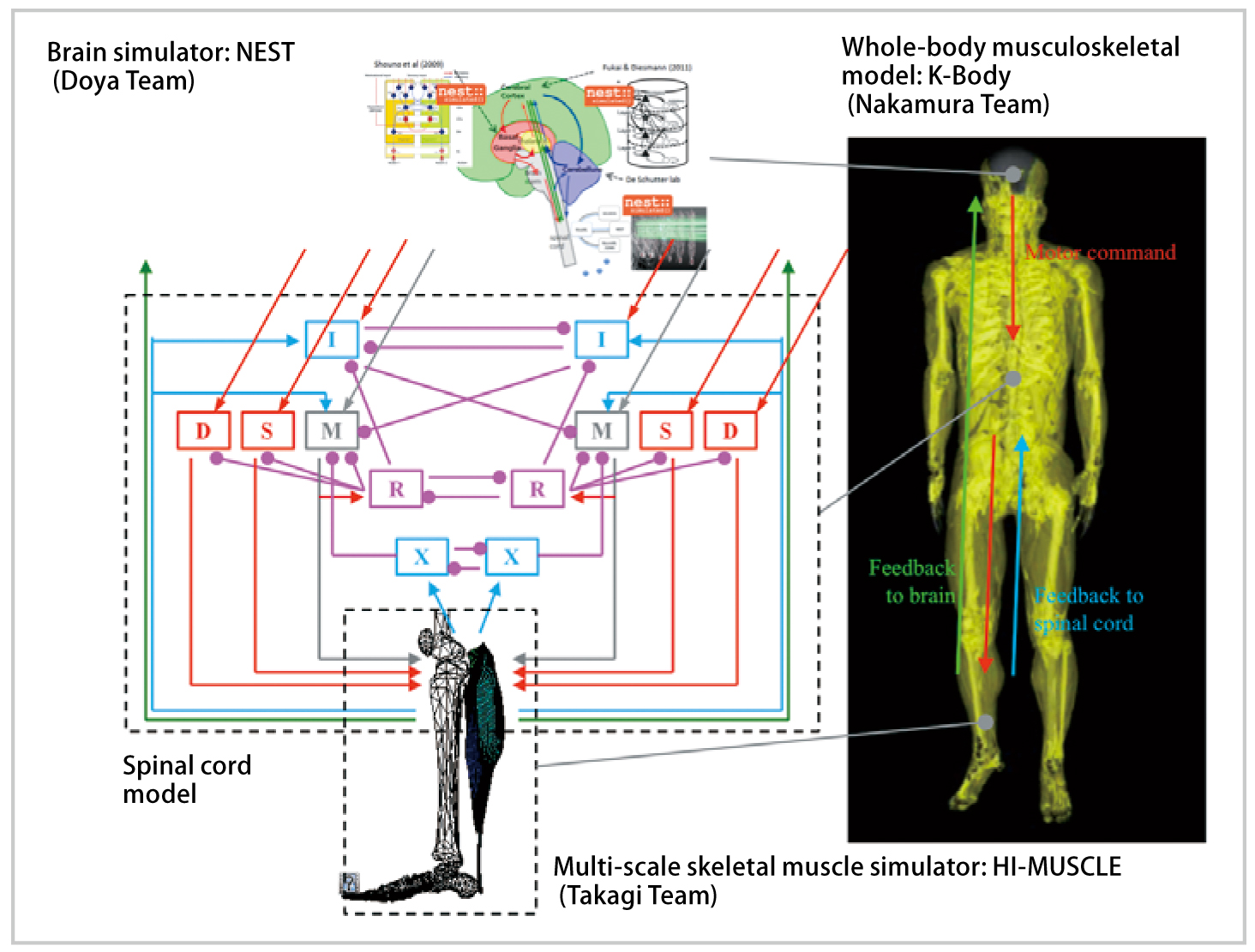
We will aim to reproduce Parkinson`s disease, which is one of the motor dysfunction resulting from a cranial nerve disease, by integrating the brain nervous system simulator and the musculoskeletal simulator.
BioSupercomputing Newsletter Vol.8
- SPECIAL INTERVIEW
- Grand Challenge opens the way to the future of life science through innovative approach
Program Director, Computational Science Research Program, RIKEN
Koji Kaya - A Landmark Project that Brought on an Innovation in the Field of Life Scienc
Deputy-Program Director, Computational Science Research Program, RIKEN
Ryutaro Himeno
- Report on Research
- Multi-scale, multi-physics heart simulator UT-Hear
Graduate School of Frontier Sciences, the University of Tokyo
Toshiaki Hisada, Seiryo Sugiura, Takumi Washio,
Jun-ichi Okada, Akihito Takahashi
(Organ and Body Scale WG) - Simulation Model for Insulin Granule Kinetics in Pancreatic Beta Cells
Graduate School of System Informatics, Kobe University
Hisashi Tamaki(Cell Scale WG) - The road to brain-scale simulations on K
Brain Science Institute, RIKEN, Institute of
Neuroscience and Medicine (INM-6),
Juelich Research Center
Medical Faculty, RWTH Aachen University Markus Diesmann
(Brain and Neural Systems WG) - MD Core Program Development for Large-scale Parallelization
Computational Science Research Program, RIKEN
Yousuke Ohno(High-performance Computing Team)
- SPECIAL INTERVIEW
- Aiming to realize hierarchical integrated simulation in the circulatory organ system
and the musculoskeletal / cerebral nervous systems
Professor, Department of Mechanical Engineering and Department of Bioengineering The University of Tokyo
Shu Takagi(Theme3 GL) - Leading-edge large-scale sequence data analysis with K computer in order to promote the understanding of life programs and their diversity
Professor, Human Genome Center, The Institute of Medical Science, The University of Tokyo
Satoru Miyano(Theme4 GL)
- Report
- 4th Biosupercomputing Symposium Report
Computational Science Research Program, RIKEN
Eietsu Tamura - “K Computer” Compatible Computer: Installation of SCLS Computer System
HPCI Program for Computational Life Sciences, RIKEN
Yoshiyuki Kido
