
Looking back on Six and a Half Years of the“ Next-Generation
Integrated Simulation of Living Matter”
Grand Challenge opens the way to the future
of life science through innovative approach

Program Director, Computational Science Research Program, RIKEN
Koji Kaya
●Six-and-a-half-year support of the “K computer” from the viewpoints of hardware and software
When RIKEN took the central role in the development of today’s “K computer” and worked with many public research institutions and universities in its development, and made environmental arrangements for its usage, we carried out many discussions at RIKEN Science Council which I chaired at that time. Although studies utilizing supercomputers had progressed across a wide range of areas including material science, high end computation did not play an important role in the life science field. In these circumstances, RIKEN ended up making the proposal that supercomputer-based studies would become very important in the life science field, and RIKEN should take a proactive stance. I think administration officers including the Ministry of Education, Culture, Sports, Science & Technology which took the initiative in the project probably held active discussions. However, I believe that our proposal contributed to appointment of RIKEN as the core agency.
Along with developing a supercomputer, the R&D of Grand Challenge Application software to allow maximum utilization of the new computer became one of the major goals of the project. Regarding “Next-Generation Integrated Nanoscience Simulation Software,” a supercomputer had already been introduced in the early 2000s when I was Director of the Institute for Molecular Science, National Institutes of Natural Sciences. Then, theorists involved in nano-technology gathered to start joint study of computational science. Therefore, the Institute for Molecular Science served as a center to start the development of Grand Challenge simulation software for understanding and predicting the characteristics and phenomena of nanomaterials through electronic-, atomic- or molecular-level sophisticated, large-scale computation. However, regarding the life science field, the computational science approach had hardly been explored. Still, Dr. Ryutaro Himeno and Dr. Makoto Taiji et al insisted that research and development should be promoted in this field, and played a driving force for Grand Challenge “Next-Generation Integrated Simulation of Living Matter Software.” As their insight was brilliant and the introduction of computational science in life science was a must, 13 institutes (ultimately 15 institutes) gathered to establish the R&D system. It officially started in September 2006 after its application was approved by the Ministry of Education, Culture, Sports, Science & Technology in June 2006. Since I did not major in life science, I honestly hesitated for a moment when I was asked to assume the position of Program Director. However as I was once involved in computational science in the field of nanotechnology, I thought I could be of some help in organizational control and decided to assume the position. From 2008, I have been concurrently serving as Deputy Director of the RIKEN Next-Generation Supercomputer R&D Center. As a result, I have watched over the development of the “K computer” from the viewpoints of both hardware and software.
●Tangible results produced by Grand Challenge
While no one in the material science field had doubts about the research and development of simulation software, it was initially very difficult to gain understanding from those involved in the life science field. After all, we had to address a considerably broad spectrum ranging from the molecular level to whole-body scale. What we had to do was a multi-scale simulation. In addition, we had hardly any established theory for each hierarchy. Regarding cells for example, RIKEN of course had been studying them by experiments. However, cells are not simple, anyway. The cell contains various substances including proteins, whose concentrations in the cell are incredibly high. It is almost unbelievable that proteins can be dissolved in water at such high concentrations. Everyone doubted whether they could replicate it correctly and calculate cell functions. Cell simulation was such a very difficult theme. However, as a result, research has progressed in those 6 years so that we can see some daylight. Of course it is also a fact that actual replication of the cell is far beyond the computational capability of the “K computer.”
After I had just assumed the position of Program Director of Grand Challenge, our group including Dr. Himeno met many leading professors in the life science field. Many of them gave us rather dismissive answers. They neither showed active interest in nor extended support to our attempt. Only 6 to 7 years ago, it was a trend in the life science field. Now our research and development in life science has advanced by leaps and bounds. Simulation studies have advanced understanding at a cellular level, and also at the molecular level we can now replicate various protein functions. Brain research has advanced such that the “K computer” would in a real sense replicate functions like those of a real brain on a computer for the first time in the world. The Organ and Body Scale WG has enabled very excellent heart simulation. In blood flow simulation, a radically new fluid-structure interaction analysis method was developed for replication of the thrombogenic process. Sonic simulation studies which may lead to the development of ultrasonic therapy apparatus also show successful results. These are very important achievements also in terms of the contribution to medical care and medical engineering. In the field of data analysis fusion, some are initially doubtful about processing bioinformatics data by a petaflops-class supercomputer. When the environment necessary for easier genome analysis is put in place, the era of personalized medicine which requires the processing of enormous amounts of data will arrive. In this context, the supercomputer with high calculation performance was proven to be useful in various processing tasks.
Initially, many people doubted whether a computational science-based approach can provide adequate results in the complex and broad-ranging life-science field. In spite of this background, there is no doubt that we got some positive response and discovered many possibilities during those 6 to 7 years. Until we reached this stage, the High-performance Computing Team led by Dr. Taiji took a key role in the development of software technologies and supported this project. I cannot forget their important role. Of course our accomplishments are still a far cry from clarification of complex l ife phenomena and mechanisms. However, it is a fact that tangible results were obtained. I think Grand Challenge made a powerful impact. I do feel that high end computation will make a strong positive impact on the biology and medical care in the 21st century.
 |
 |
Building of Advanced Institute for Computational Science under construction (April, 2009) |
Supercomputer room under construction was shown to the press (February, 2010) |
 |
 |
Commemorative picture for completion of“ K computer” |
The 3rd BioSupercomputing Symposium (March, 2011) |
●Hoping for young people’s contribution to development of future life science
Another achievement of Grand Challenge is that as simulation science grew in the field of life science, many young researchers became interested and joined the field, and we can now collaborate with people from a wide range of research fields. I believe those young scientists will become a very important human resource which supports future research. However, the problem is that the new computational science, which those young researchers will be engaged in, has not yet won enough recognition in Japan. In the USA where computer science enjoys greater recognition, reasonable posts are offered to researchers engaged in computational science, but the Japanese life science field doesn’t even go that far. Still today, vacant posts tend to be allocated to experimental scientists. Simulation studies will definitely have a key role in future life science. However, I cannot help feeling that professors who have been leading life science researchers do not seem so sure about that. In addition, we have to make the best of today’s tight research budget. I think this is one of reasons why it is not easy to increase the number of posts for the computational bio-science.
The Grand Challenge Program ends in March 2013. While producing excellent results, there still remain many courses of research and development which have to be challenged again by the collaborations among colleagues in the Grand Challenge Program. I believe how to inherit and expand it will be a major challenge from now on. The life science field had a very limited interaction and collaboration with other research fields. However, in Grand Challenge, researchers across various fields gathered to create a new life science. I hope more and more young researchers overcome the barrier, absorb the knowledge and information in various fields, and pave the way for a new world.
In any age, it is always hard to open up a new field, and we may face many difficulties in doing so. However, one thing that I can say without fear is that life science is facing an era of revolutionary change. More and more innovations must be produced from now. I believe Grand Challenge is a runway for that. It is my fervent hope that as many young researchers as possible take off from this runway, and continue the effort to blaze a new path to the future.
 |
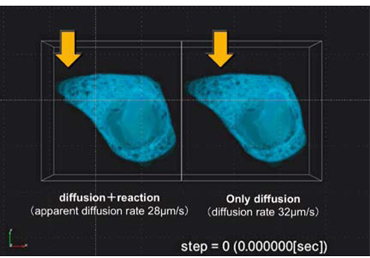 |
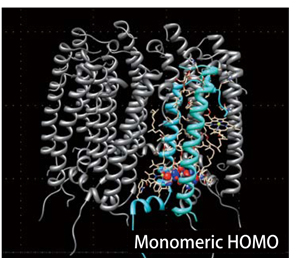
Protein DF - Protein All Electron Calculation |
Ca+ kinetics simulation(HepG2) |
BioSupercomputing Newsletter Vol.8
- SPECIAL INTERVIEW
- Grand Challenge opens the way to the future of life science through innovative approach
Program Director, Computational Science Research Program, RIKEN
Koji Kaya - A Landmark Project that Brought on an Innovation in the Field of Life Scienc
Deputy-Program Director, Computational Science Research Program, RIKEN
Ryutaro Himeno
- Report on Research
- Multi-scale, multi-physics heart simulator UT-Hear
Graduate School of Frontier Sciences, the University of Tokyo
Toshiaki Hisada, Seiryo Sugiura, Takumi Washio,
Jun-ichi Okada, Akihito Takahashi
(Organ and Body Scale WG) - Simulation Model for Insulin Granule Kinetics in Pancreatic Beta Cells
Graduate School of System Informatics, Kobe University
Hisashi Tamaki(Cell Scale WG) - The road to brain-scale simulations on K
Brain Science Institute, RIKEN, Institute of
Neuroscience and Medicine (INM-6),
Juelich Research Center
Medical Faculty, RWTH Aachen University Markus Diesmann
(Brain and Neural Systems WG) - MD Core Program Development for Large-scale Parallelization
Computational Science Research Program, RIKEN
Yousuke Ohno(High-performance Computing Team)
- SPECIAL INTERVIEW
- Aiming to realize hierarchical integrated simulation in the circulatory organ system
and the musculoskeletal / cerebral nervous systems
Professor, Department of Mechanical Engineering and Department of Bioengineering The University of Tokyo
Shu Takagi(Theme3 GL) - Leading-edge large-scale sequence data analysis with K computer in order to promote the understanding of life programs and their diversity
Professor, Human Genome Center, The Institute of Medical Science, The University of Tokyo
Satoru Miyano(Theme4 GL)
- Report
- 4th Biosupercomputing Symposium Report
Computational Science Research Program, RIKEN
Eietsu Tamura - “K Computer” Compatible Computer: Installation of SCLS Computer System
HPCI Program for Computational Life Sciences, RIKEN
Yoshiyuki Kido
