
Looking back on Six and a Half Years of the“ Next-Generation
Integrated Simulation of Living Matter”
A Landmark Project that Brought on
an Innovation in the Field of Life Science

Deputy-Program Director, Computational Science Research Program, RIKEN
Ryutaro Himeno
●Phenomenal Findings by the Project
The “Next-Generation Integrated Simulation of Living Matter,” a research and development project which we have committed to since October of 2006, essentially aimed to develop a “show-case model” of simulation software in the life science field as part of the “Next-Generation Super Computer Project”. This project was set up to prove that “new and astounding studies could be performed” with a supercomputer with a performance at the 10 PFLOPS level.
Looking back on the last six and a half years, there were many things we could and could not accomplish. But in the perspective of research and development of simulation software, I believe we were able to produce a large number of software streamlined for the “K computer”, which were able to display a higher performance than the theoretical performance that we originally expected.
Not only did the “MD core program for large-scale parallel computers (cppmd)”, the molecular dynamics software streamlined for the “K computer”, result in a performance value close to 40% as expected, the “Fluid-structure interaction analysis program for whole body voxel simulation (ZZ-EFSI)” also attained a remarkable performance value of 43%. Further, the “multi-scale multi-physics heart simulation (UT-Heart)” also attained a performance value close to 30%, and “non-invasive treatment simulation (HIFU)” also attained a performance value higher than 20%. Originally, our goal was to develop simulation software that exceeded a performance of 1 PFLOPS using the entire “K computer”, and we were hoping to possibly develop two software. But these four software will definitely result in performance values higher than 20% if they are actually run on the entire “K computer”. Therefore, it might sound a little overstated, but I feel we can say we surpassed our initial goal. Regardless of how we say it, simulation software that yields high performance values has been created. There are thirteen software including these four that were able to produce solid results on ten thousand nodes of the “K computer”. The fact that one third of approximately thirty software programs in total attained honorable records is a success that surpasses our original expectations.
You may think, “Isn’t that a little over dramatic for 30%?”, but in the year 2006 when the project started, there was much speculation and criticism regarding development of simulation software in the life science field for the supercomputer. In those terms, to be able to develop thirteen usable software is a wonderfully remarkable achievement.
Looking back, it was good that we were able to produce respectable findings every year. People have gradually started to accredit our accomplishments through these yearly achievements. Most of all, the actual researchers who are developing the software understand that their efforts for improving parallel performance have been fed back to them as achievements, and are at a stage where they say, “We can’t operate without the “K computer”, actually even the “K computer” isn’t enough”. It is now hard to believe that there were worries like, “Can we fully utilize a 10 PFLOPS machine?” when we first started.
Especially in the year 2012, we only had a period of two months starting in July to be granted priority to use 10 PFLOPS together with the five fields of the Strategic Programs for Innovative Research. We had a difficult time on how to handle everyone’s concerns regarding the lack of calculation time. Reflecting on our initial worries, we can say it is a gratifying miscalculation, but it is very disappointing that some were not able to produce sufficient scientific findings due to lack of time to use on a priority basis. For example, the simulation software “Neural Simulation Tool (NEST)”, which replicates input and output of the entire brain in order to reproduce the growth and learning capabilities of the brain, has the largest calculation in the field of brain science in the world when it uses the entire “K computer”, but we could not get enough calculation time. In that context, I believe a lot more could be accomplished if we had another year.
Likewise, there are many topics that are left uncompleted, but I feel that the project was able to secure respectable achievements overall. Without this project, I believe there still would have been many researchers that would say “the “K computer” is not necessary for the field of life sciences”.
 Ryutaro Himeno, Deputy-Program Director, who gave his lecture at “Next- Generation Integrated Simulation of Living Matter Symposium 2007” |

|
 |

|
Poster Session held at the Joint Computational |
Poster Session held at the 2nd Biosupercomputing Symposium (March, 2010) |
●Integration Born from Summer School
The “K computer” has a relatively fast network and is constructed with highly distinguished hardware. The work of the High-performance Computing Team whose researchers found bugs in compiler before others use it and made various efforts to bring out full performance values cannot be ignored. Their work is wonderful. Through their steady efforts, the “K computer” has been built as a machine that can be used with ease, and I believe the existence of this team has led to the development of a large number of software that attain high efficiency. For improving computation speed, it is critical to share knowhow among all software development researchers. The role they played in sharing a precise pool of information for generating higher performance was exceptional.
The word “sharing” brings to mind the early stage of this project. Personnel with research backgrounds of various scales from many research institutions and universities joined the project, but since they spoke their own technical language and abbreviations, it was difficult to understand each other when we initially held symposiums (laughing). Although explanations became more explicit and understandings deepened, it did not lead to integration. In the midst of this, the summer school in which we collected younger researchers was effective. While each researcher worked on his/her new model, people from varying research fields used the computers in a similar manner but with a different approach, and this observation directly stimulated everybody. Through summer school, researchers became to understand what researchers from other fields were pursuing, which led to the overcoming of research field and technical language barriers, and eventually progressed to integration. You can say that the simulation of blood clotting through the fluid-structure interaction analysis method was also born out of this new kind of integration.
I am very pleased that this project stimulated growth in each field due to the new integration. This itself cannot be explained in a research paper, and may not be perceived as an achievement. Even so, I hope there will be a day when people will say that in the long run this project became the turning point for generating new connections, and led to wide-ranging progress in research and development.
●To Expand on the Accomplishments of the Project
To simulate the phenomenon of life which is a multi-body and multi-layered system, it is necessary to explore new phases through new integrations. In this project, new integrations have been formed in each research and development team, and interesting findings are coming to light. The main focus of the project was software development, but at this point interests are shifting towards the research findings generated from the new software which is an exciting result. You can also say we were science-driven and software had been developed as seedlings for the researchers’ interests, but some could possibly be put to practical use for actual drug discovery and medical care.
The project will end in the end of fiscal 2012, and each research and development team will be dissolved, but those that have the potential for future needs will be incorporated into the Strategic Programs for Innovative Research (field 1). Other software that should be inherited and extended, rather than ending up as they are, will be undertaken by RIKEN’s Advanced Center for Computing and Communication for further development, updates, and support. We hope they will be actually used by the research centers at RIKEN. We will need to maintain a running environment, or else efforts will end with the development stage. By continuing the efforts in RIKEN’s research topics, I hope many will realize and appreciate the value of operating this project at RIKEN.
I mentioned earlier, the young researchers whose research attitudes have changed through the summer school. We have founded a bio-supercomputing research group in order to support them, and hope to continue to help this research group evolve. It will be a wonderful place to continue to cultivate these budding young researchers that possess new insights.
 |
 |
Ultrasound treatment simulator: ZZ-HIFU |
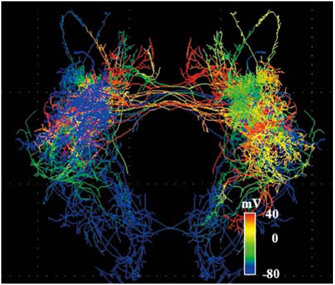
Neural network activity simulation |
BioSupercomputing Newsletter Vol.8
- SPECIAL INTERVIEW
- Grand Challenge opens the way to the future of life science through innovative approach
Program Director, Computational Science Research Program, RIKEN
Koji Kaya - A Landmark Project that Brought on an Innovation in the Field of Life Scienc
Deputy-Program Director, Computational Science Research Program, RIKEN
Ryutaro Himeno
- Report on Research
- Multi-scale, multi-physics heart simulator UT-Hear
Graduate School of Frontier Sciences, the University of Tokyo
Toshiaki Hisada, Seiryo Sugiura, Takumi Washio,
Jun-ichi Okada, Akihito Takahashi
(Organ and Body Scale WG) - Simulation Model for Insulin Granule Kinetics in Pancreatic Beta Cells
Graduate School of System Informatics, Kobe University
Hisashi Tamaki(Cell Scale WG) - The road to brain-scale simulations on K
Brain Science Institute, RIKEN, Institute of
Neuroscience and Medicine (INM-6),
Juelich Research Center
Medical Faculty, RWTH Aachen University Markus Diesmann
(Brain and Neural Systems WG) - MD Core Program Development for Large-scale Parallelization
Computational Science Research Program, RIKEN
Yousuke Ohno(High-performance Computing Team)
- SPECIAL INTERVIEW
- Aiming to realize hierarchical integrated simulation in the circulatory organ system
and the musculoskeletal / cerebral nervous systems
Professor, Department of Mechanical Engineering and Department of Bioengineering The University of Tokyo
Shu Takagi(Theme3 GL) - Leading-edge large-scale sequence data analysis with K computer in order to promote the understanding of life programs and their diversity
Professor, Human Genome Center, The Institute of Medical Science, The University of Tokyo
Satoru Miyano(Theme4 GL)
- Report
- 4th Biosupercomputing Symposium Report
Computational Science Research Program, RIKEN
Eietsu Tamura - “K Computer” Compatible Computer: Installation of SCLS Computer System
HPCI Program for Computational Life Sciences, RIKEN
Yoshiyuki Kido
