
Biosupercomputing Pioneers the Future of Life Science
The time has come for biosupercomputing to get results with the world's No. 1 supercomputer “K computer”, and take up the challenge of "prognostic biology".

Deputy Program Director of Computational Science Research Program, RIKEN
Ryutaro Himeno
●One third of the software has started tests in the environment of “K computer”.
In 2006, the grand challenge (ISLiM:Next- Generation Integrated Simulation of Living Matter) started with the mission of understanding the super n-body multi-level problem, which is an extremely complex life phenomenon, using K computer's peta-scale computation ability with unprecedented performance, and develop software that will contribute to drug discovery and medical care. Five years has passed, and software development has now reached a new phase to proceed with super-large scale parallelization and machine tuning using “K computer”.
According to the 37th TOP500 announced June of this year (2011), “K computer” has measured 8.162 PFLOPS on the LINPACK benchmark (execution efficiency of 93.0%), and achieved 1st place even though it was still under construction. Currently, our 31 software projects that are to run on this highly acclaimed “K computer” are under development. 11 of these have already passed the development phase for large scale parallelization (8,192 parallel), and are being tested on “K computer”. The "MD core program for large-scale parallel computers (cppmd)" is one of them, has an execution efficiency that exceeds 30%, and reported 1.3PFLOPS as of the end of March. It is required for these software developments to make as much use of the performance of "K computer" as possible. We want to have a high line-up of software that exceeds at least 1PFLOPS when using the whole machine (10PFLOPS), and as for the mean time, one of the softwares has already achieved the target performance. Including cppmd, four softwares (cppmd, ZZ-EFSI, CafeMol, ZZ-HIFU) have exceeded an execution efficiency of 20%, and two (cppmd, ZZ-EFSI) have reached approximately 40% as of now (October 2011). Keeping in mind that we were able to start working on “K computer” from the beginning of this year, performance tuning has progressed at a rapid pace, and performance results are rather satisfactory. However, further improvements are under way in some softwares which have poor single performance values or have good parallel performance but do not have good single performance. Of course, “K computer” itself is a machine still under construction, and compilers and network libraries will be enhanced. These enhancements will probably improve the software performance, but we have no time to waste. The time allocated to us to have priority access to “K computer” is until the end of October, 2012, when its shared services will start. Until then, we must organize calculations and produce results that have a strong scientific value. Therefore, we need to make our best efforts, such as optimizing code, to withdraw the highest performance in the shortest
amount of time.
We of course are planning to run on “K computer” all of the 31 softwares that are being developed and get results. But since time is limited, we will have to decide on a detailed plan for how to allocate calculation size and time. There may be 10, or maybe even less, that can be tested using the entire system. Furthermore, we need to debate on the setup, such as reducing calculation size but spending a longer time. One big challenge will be devising a way to maximize our accomplishments by utilizing calculation resources in a limited time.
●Grand challenge sows seeds for opening doors to the new generation of life science
In parallel to the grand challenge, the "HPCI STRATEGIC PROGRAM Supercomputational Life Science", started this year. The main mission of the grand challenge is software development which is science-oriented, and not needs-oriented. With our desire to lead the entire life science field towards computer simulation and data analysis, we have created software packages for a wide variety of research fields, such as genetics, biomolecules, organs, and also the whole body. By producing further results and publicizing them, our goal is to lower the threshold of this genre, and thus allow it to be used by people who wish to pursue new research. Therefore, our software lineup does not necessarily consist of software that may be desired by the pharmaceutical and medical industries. On the other hand, for the HPCI Strategic Program, we will maximize the use of high computing resources such as “K computer”, and aim to produce results that will have social value. Therefore, taking the same computational software for molecular dynamics as an example, development on “K computer” is its mission to meet the grand challenge, and as for the Strategic Program, it is expected to get results that will be beneficial to drug discovery. Like "development" and "usage", its use may be entirely different even if the same topic is addressed. It is as though we have planted seeds and are striving our best for beautiful flowers to bloom. With this analogy, we can say it is the job of the Strategic Program to nurture it until it bears fruit, and then to harvest it.
We, in a sense, have been running towards a goal that has been set up without considering the needs of society. However from now on, it is also expected for the project to be applied for actual drug discovery and for development of medical equipment, and it will be required to add new functions, work along those who want use it for collaborative research, and to convene lectures. Through these activities, I believe the results of the grand challenge will be carried on with the Strategic Program. There already are high hopes in the findings by the molecule-scale research team and the data analysis integration research team, especially in the drug discovery field. Attention has been drawn to the fact that this software can already be useful. It may take some more time for development by the Organ and Body Team, but they too are also expected to contribute to drug discovery and medical care. I hope that by adding new functions and combining them based on the software, the software will grow in unpredictable ways. But to begin with, what we must first do is to enhance and perfect the software. With these efforts, there is no doubt we will achieve research results that will surprise the world, namely, the blossoming of a beautiful flower of science.

An array of the quadrillion computer “K computer” in the computer room (left) .
One chassis is equipped with 24 system boards (center).
Building of the Advanced Institute for Computational Science in which “K computer” is being installed (right).
●Being able to use “K computer” has strengthened a sense of unity between researchers
When the grand challenge was in blueprint, an overwhelming majority of people around us felt that, in order to accelerate life science research, enhancing experimental devices would be much more effective than spending money on supercomputer. The grand challenge started with voiced doubts as to whether life science software could produce decent results using a supercomputer. Already at this point, however, one third of the software being developed had started testing on “K computer”, and produced results that exceed 10 thousand parallels. The number of softwares with 10% or higher execution efficiency is rising. Of course, this is only stating the performance values for the software program, and accomplishments that have scientific value will be coming henceforth. But we can say for certain that the opinions of people concerned with this project are changing. Expectations in the possibilities of supercomputing are growing, thick and fast.
At the same time, the project members have also changed. Since the members came from a wide range of research fields, many only had interests in their own field, and in the beginning when we started where there were no common topics to discuss. Facing the same table, we were able to identify each other’s faces and were able to have some understanding among one another, but a dramatic change occurred when “K computer” became available to us. “What can be done in order to achieve high performance with “K computer”?” This question is shared by every researcher involved in the project. The sense that they are together in driving the same project forward with “K computer” at its heart has been born through shared experiences and a common language, and the bond between each member has strengthened. One example can be seen in the mailing list recently founded for young researchers working on “K computer”. Passionate messages can be seen there, such as, "“K computer” is now the world best's in hardware, next it's for us to achieve something with it", and "it's up to us to try our best". Once again, it was made clear to me that the project is moving forward with everyone having a high sense of purpose and togetherness.
Our mission is to produce results at the same time “K computer” goes into action. The time left for us is only one more year. The final year should be the most important to make a beautiful flower blossom, and go on to the next phase, "usage". In getting ready for the prospective year, the biggest concern is next year's budget. To secure our budget is also a major challenge.
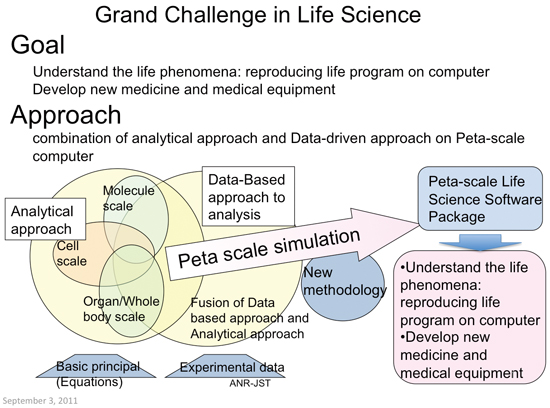
Aiming at development of software for contributing to the understanding of life phenomena and medical care by utilizing the full performance of the "K computer".
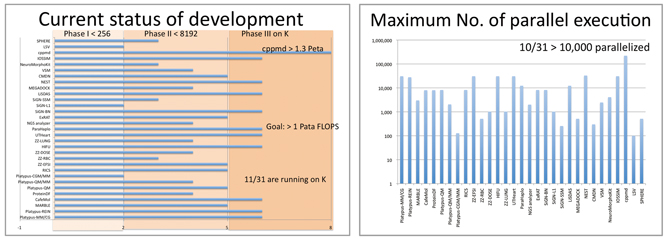
Of 31 kinds of software under development, 11 achieved the scheduled massive parallelization and are being tested with “K computer” (left). There are 10 kinds of software which have achieved 10 thousands parallelization or more (right) (as of October 2011).
BioSupercomputing Newsletter Vol.5
- SPECIAL INTERVIEW
- The time has come for biosupercomputing to get results with the world's No. 1 supercomputer
"K computer", and take up the challenge of "prognostic biology".
Deputy Program Director of Computational Science Research Program, RIKEN Ryutaro Himeno - What should we do to promote industrial use of sophisticated computer resources and development applications?
Chief Coordinator of Foundation for Computational Science Masahiro Fukuda
Chief Researcher of Urban Innovation Institute and Executive Board Member and Bureau Chief of BioGrid Center Kansai Ryuichi Shimizu
- Report on Research
- Analysis of molecular mechanism of enzymatic reactions by QM/MM Free Energy Method
Graduate School of Science, Kyoto University Shigehiko Hayashi (Molecular Scale WG) - Computational Mechanobiology of Actin Cytoskeleton
Institute for Frontier Medical Science, Kyoto University Yasuhiro Inoue (Cell Scale WG) - Development of Blood flow Analysis Method for Simulation of Thrombus Formation
Department of Mechanical Engineering, The University of Tokyo Satoshi Ii (Organ and Body Scale WG) - Development of Data Assimilation Technology for Simulation of Living Things
The Institute of Statistical Mathematics Tomoyuki Higuchi (Data Analysis Fusion WG)
- SPECIAL INTERVIEW
- Pioneering the Future of Computational Life Science toward Understanding and Prediction of Complex Life Phenomena
Program Director of RIKEN HPCI Program for Computational Life Sciences Toshio Yanagida
Deputy- Program Director of RIKEN HPCI Program for Computational Life Sciences Akinori Kidera
Deputy- Program Director of RIKEN HPCI Program for Computational Life Sciences Yukihiro Eguchi
- Report on Research
- Simulation Applicable to Drug Design
Research Center for Advanced Science and Technology, The University of Tokyo Hideaki Fujitani (Field 1- Program 2) - An Ultra-fast Analysis System for Next-Generation DNA Sequencer Data
Graduate School of Information Science and Engineering, Tokyo Institute of Technology
Yutaka Akiyama, Takashi Ishida, Masanori Kakuta, Shuji Suzuki (Field 1- Program 4)
- Report
- BioSupercomputing Summer School 2011
Computational Science Research Program, RIKEN Yasuhiro Ishimine (Organ and Body Scale WG)
Research and Development Center for Data Assimilation, Institute of Statistical Mathematics Masaya Saito (Data Analysis Fusion WG)
Niigata University of International and Information Studies Eisuke Chikayama (Cell Scale WG)
School of Medicine, Tokai University Yohei Nanazawa (Cell Scale/Organ and Body Scale WG)
Computational Science Research Program, RIKEN Takashi Handa (Brain and Neural Systems WG)
Computational Science Research Program, RIKEN Gen Masumoto (High-performance Computing Team)
Computational Science Research Program, RIKEN Kei Moritsugu (Molecular Scale WG) - “Next-Generation Integrated Simulation of Living Matter (ISLiM)”, a web page dedicated to new applications, has opened.
Computational Science Research Program Integrated Simulation of Living Matter Group
- Event information
