
Development of Blood flow Analysis Method for Simulation of Thrombus Formation

Department of Mechanical Engineering, The University of Tokyo
Satoshi Ii (Organ and Body Scale WG)
When the inner wall of a blood vessel is damaged, a thrombus is formed at the site of damage. Due to the thrombus, bleeding from a slight injury usually stops after a certain amount of time. This is the major role of a thrombus. The thrombus formed is lysed after bleeding stops. For some reason, it sometimes does not disappear and becomes enlarged. Then, it may block the blood vessel, or the enlarged thrombus may slough off and occlude a more distal vessel to block blood flow. Such a thrombus causes brain infarct and cardiac infarct. It is also a cause of the development of economy-class syndrome or flight-related DVT. For these reasons, antithrombosis agents are used in clinical practice. However, since they have the potential to affect normal thrombus involved with hemostasis, they should be given cautiously. However it is known to be very difficult to assess the effect of a medicine in the bloodstream of the human body in advance. Against this background, our group is aiming to reproduce thrombus formation through simulation. For example, we are thinking of its use for understanding drug efficacy in the bloodstream in developing new drugs.
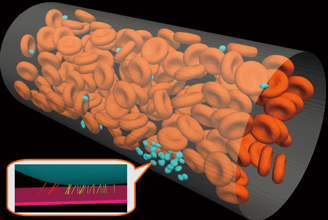
In the thrombus formation process, adhesion of platelets to the damaged vessel wall, agglutination of platelets and coagulation incorporating erythrocytes, involving various factors such as the bloodstream and cell metabolism, occur in a step-by-step manner. In this research, we focus on the early process, platelet primary aggregation in which platelets adhere to the damaged cell wall. In this process, platelets adhere to the damaged cell wall through binding of vWF (von Willerbrand factor) attached to collagen beneath vascular endothelial cells and GPIbα (glycoprotein Ibα) found on the platelet surface. In order to analyze such phenomena, we are developing an approach to deal with bloodstream phenomena including erythrocytes and platelets, and GPIbα-vWF binding phenomena between the platelet surface and damaged cell wall in a unified manner (Fig.1).
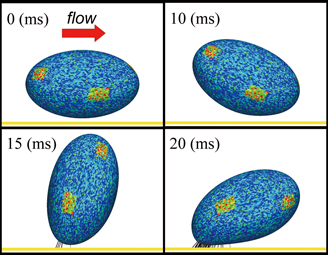
Since erythrocytes have a very high bulk ratio of around 45% in the bloodstream, when the vessel diameter is micrometer-size, they affect the flow properties of the blood stream and become strongly involved in platelet aggregation. Since erythrocytes are very flexible and deformable, they cause an interacting phenomenon with the flow field. Since these phenomena must be analyzed in a system containing numerous erythrocytes, our group proposes a new analytical approach which excels in calculation efficiency and has a good compatibility with blood flow analysis [1,2]. We performed an analysis by use of an experimentallyobtained physical property of erythrocytes. As shown in Fig 2, we obtained a valid result as compared to the deformation behavior of real erythrocytes in the flow field.
In the analysis of platelet primary aggregation, binding of GPIbα on the platelet surface and vWF in the damaged cell wall has to be considered. They are molecular-scale phenomena, but their interactive analysis in combination with blood flow is currently impossible. Therefore, we introduced a technique in which we can deal with such binding phenomena statistically, and combined it with blood flow analysis. Fig 3 shows an example of the analysis. It reproduces how binding of GPIbα on the platelet and vWF at the bottom of the wall (simulating a damaged vessel wall) takes place, and how platelets adhere to the cell wall.
Hereafter, we will deal with aggregation of platelets by modeling the metabolic response of platelets, and reproduce the thrombus formation process in a step-by-step manner.
【References】
[1] K. Sugiyama, S. Ii, S. Takeuchi, S. Takagi and Y. Matsumoto, A full Eulerian finite difference approach for solving fluid-structure coupling problems, J. Comput. Phys., 230 (2011) 596-627.
[2] S. Ii, X. Gong, K. Sugiyama, J. Wu, H. Huang and S. Takagi, A full Eulerian fluid-membrane coupling method with a smoothed volume-of-fluid approach, Commun. Comput. Phys. (2011) accepted.
[3] P. Gaehtgens, C. Dührssen and K.H. Albrecht, Motion, deformation, and interaction of blood cells and plasma during flow through narrow capillary tubes. Blood Cells, 6 (1980) 799-817.
 |
|
Fig 1 : View showing a frame format of analysis toward platelet primary aggregation |
Fig. 2 : Deformation behavior of erythrocytes in the flow field [3], and result of simulation |
 |
|
Fig 3 : How platelets adhere in fluid analysis taking GPIbα-vWF binding into account |
BioSupercomputing Newsletter Vol.5
- SPECIAL INTERVIEW
- The time has come for biosupercomputing to get results with the world's No. 1 supercomputer
"K computer", and take up the challenge of "prognostic biology".
Deputy Program Director of Computational Science Research Program, RIKEN Ryutaro Himeno - What should we do to promote industrial use of sophisticated computer resources and development applications?
Chief Coordinator of Foundation for Computational Science Masahiro Fukuda
Chief Researcher of Urban Innovation Institute and Executive Board Member and Bureau Chief of BioGrid Center Kansai Ryuichi Shimizu
- Report on Research
- Analysis of molecular mechanism of enzymatic reactions by QM/MM Free Energy Method
Graduate School of Science, Kyoto University Shigehiko Hayashi (Molecular Scale WG) - Computational Mechanobiology of Actin Cytoskeleton
Institute for Frontier Medical Science, Kyoto University Yasuhiro Inoue (Cell Scale WG) - Development of Blood flow Analysis Method for Simulation of Thrombus Formation
Department of Mechanical Engineering, The University of Tokyo Satoshi Ii (Organ and Body Scale WG) - Development of Data Assimilation Technology for Simulation of Living Things
The Institute of Statistical Mathematics Tomoyuki Higuchi (Data Analysis Fusion WG)
- SPECIAL INTERVIEW
- Pioneering the Future of Computational Life Science toward Understanding and Prediction of Complex Life Phenomena
Program Director of RIKEN HPCI Program for Computational Life Sciences Toshio Yanagida
Deputy- Program Director of RIKEN HPCI Program for Computational Life Sciences Akinori Kidera
Deputy- Program Director of RIKEN HPCI Program for Computational Life Sciences Yukihiro Eguchi
- Report on Research
- Simulation Applicable to Drug Design
Research Center for Advanced Science and Technology, The University of Tokyo Hideaki Fujitani (Field 1- Program 2) - An Ultra-fast Analysis System for Next-Generation DNA Sequencer Data
Graduate School of Information Science and Engineering, Tokyo Institute of Technology
Yutaka Akiyama, Takashi Ishida, Masanori Kakuta, Shuji Suzuki (Field 1- Program 4)
- Report
- BioSupercomputing Summer School 2011
Computational Science Research Program, RIKEN Yasuhiro Ishimine (Organ and Body Scale WG)
Research and Development Center for Data Assimilation, Institute of Statistical Mathematics Masaya Saito (Data Analysis Fusion WG)
Niigata University of International and Information Studies Eisuke Chikayama (Cell Scale WG)
School of Medicine, Tokai University Yohei Nanazawa (Cell Scale/Organ and Body Scale WG)
Computational Science Research Program, RIKEN Takashi Handa (Brain and Neural Systems WG)
Computational Science Research Program, RIKEN Gen Masumoto (High-performance Computing Team)
Computational Science Research Program, RIKEN Kei Moritsugu (Molecular Scale WG) - “Next-Generation Integrated Simulation of Living Matter (ISLiM)”, a web page dedicated to new applications, has opened.
Computational Science Research Program Integrated Simulation of Living Matter Group
- Event information

![流動場中の赤血球の変形挙動[3]とシミュレーション結果](images/report03_img02.jpg)