
Cell Scale WG
Cell simulation considering time-space

Computational Science Research Program, RIKEN
Yasuhiro Sunaga
All living things consist of units called cells. More than 10 million kinds
of living things including single-cell organisms such as Escherichia
coli and multicellular organisms such as humans exist on the earth,
and the functions and forms of cells are different so that they are
optimum for their living things survival. However, it is known that
living things basic living cell functions have many things in common.
A cell has a very complicated and compact structures, and in the cell,
various biochemical reactions occurs actively for each compartment
which is known as an organelle. Due to this, molecules and reactions
are differentiated to implement various living functions efficiently.
So far, much cell simulation research ignoring cell structures has been
done by grinding up organs, the aggregates of cells, to understand
the biological systems of organisms and to explore methods
of treating diseases. In the cells, life is maintained by complex
phenomena such as biochemical reactions (metabolism) and
signaling. To reproduce these biochemical reactions, many simulators
were developed, and it became possible to replicate intracellular
metabolism and signaling. However, these simulators calculate a
nondimensionalized field and replicate a chemical reaction uniformly
in the cell, a so-called 'closed bag'. In actual cells, biochemical
reactions differ for each organelle, and uneven reactions go on in the
cell depending on the intracellular transfer of substances, inside and
outside the cell, and entry and exit of substance from and to the inside and
outside of the cell. We, the Cell Scale Research and Development Team, have
been looking at these things, and are now developing the RIKEN Integrated
Cell Simulator (RICS) which aims to perfect an intracellular time-space
simulation. With this system, we undertook a coupled analysis of intracellular
biochemical reactions and substance diffusions with the equations of
reaction diffusion. This system uses the voxel analysis framework developed
by the VACD research program of RIKEN for spatial expression, which is a
simulation system that can represent the special complex space structures
of the cell.
With this system, using the cell morphology obtained from actual
microscopic data, we simulated the transporters, reactions and diffusion of
calcium ion (Ca2+) in the cell. As concerns the cell morphology, we acquired
the cell morphology and the form of the nucleus and mitochondria with
a confocal laser microscope using a HepG2 cell, human liver-derived cells
(Figure 1). The nucleus is the largest structure in the cell, and it packs DNA.
Mitochondria have the function of producing energy for cellular activity, and
are an important location for biochemical reactions such as metabolism. We
prepared three-dimensional Volume data from microscopic images to use
in the present calculation (Figure 2). We established 10 molecules involved
in buffering reactions in the cell and 24 biochemical reactions. A channel
passing Ca2+ only was localized in part of the cell membrane shown by an
arrow in Figure 2, and simulated by influx of Ca2+.
The result of simulation is shown in Figure 3. When the buffering reaction
of Ca2+ in the cell was set up (expressed as "reaction present"), the apparent
diffusion speed of Ca2+ in the cell was decreased as compared with the case
of no biochemical reaction (expressed as "reaction absent"). This suggested
that kinetics of Ca2+ close to that in actual cells can be simulated.
This RICS makes it possible to calculate the phenomena occurring in cells
such as biochemical reactions and diffusions, considering their location. In
the future, we want to model various cell functions, and to show that the
model can be utilized to elucidate the cause of drug reactions and disease.
Moreover, we want to make this system a tool that can examine a mass of
cells that functions as a tissue.
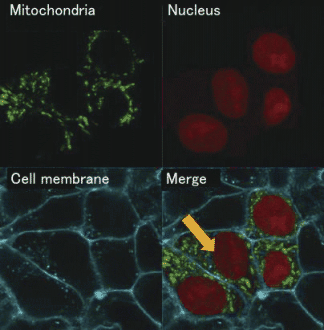
Figure 1 : Cross section image of a cell photographed using a confocal laser microscope |
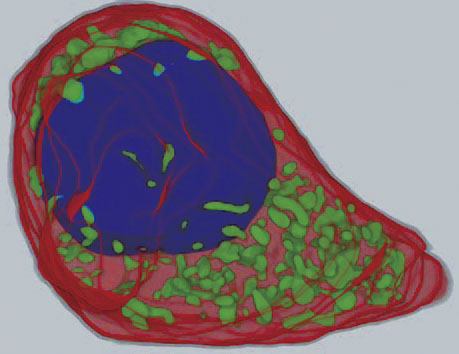
Figure 2 : Cell morphology reconstructed threedimensionally from successive cross section images of cells |
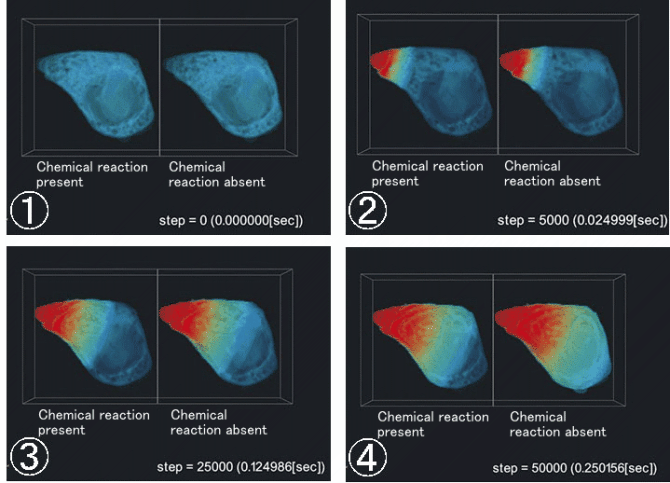
Figure 3: Time-course of Ca2+ concentrations (volume rendering)
BioSupercomputing Newsletter Vol.4
- SPECIAL INTERVIEW
- In order to change from observation-type medical practice focusing on experience to prediction-type medical practice to construct the base of theoretical medicine
Professor, Department of Internal Medicine (Cardiovascular Medicine), Director of the Metabolic Disease Research Center, Bio-Research Medical Center, Tokai University Graduate School of Medicine, and Director, Department of Metabolic System Medicine, Tokai University General Medical Laboratory Shinya Goto - It is expected that new possibilities in nutrition science and health control will be opened up
by simulation science
EXECUTIVE PROFESSIONAL Health infomatics DEPT., Ajinomoto Co., Inc. Toshihiko Ando
- Report on Research
- The functions of a multidrug discharging transporter were verified by coarse graining molecular simulation (Molecular Scale WG)
Graduate School of Science, Kyoto University Shoji Takada / Xin-Qiu Yao / Hiroo Kenzaki - Cell simulation considering time-space (Cell Scale WG)
Computational Science Research Program, RIKEN Yasuhiro Sunaga - Development of HIFU simulator for non-invasive treatment with high-intensity focused ultrasound (Organ and Body Scale WG)
VCAD System Research Program, RIKEN Kohei Okita - PLATO: Platform for a collaborative brain system modeling toward development of large scale mathematical model.(Brain and Neural WG)
①Computational Science Research Program, RIKEN
②Brain Science Institute, RIKEN
Keiichiro Inagaki①/ Takayuki Kannon②/ Nilton L. Kamiji②/ Koji Makimura②/ Shiro Usui ①②
- Report
- Report on the workshop in BMB2011 (Joint Meeting of the 33rd Congress of the Molecular Biology Society of Japan and the 83th Congress of the Japanese Biochemical Society)
- Winter School 2011 for the Integrated Simulation of Living Matter
Computational Science Research Program, RIKEN Yasuhiro Ishimine (Organ and Body Scale WG)
The Institute of Medical Science, The University of Tokyo Hidetoshi Urakubo (Brain and Neural WG)
Computational Science Research Program, RIKEN Yasuhiro Sunaga (Cell Scale WG)
Computational Science Research Program, RIKEN Gen Masumoto (High-Performance Computing Team)
Computational Science Research Program, RIKEN Keiji Misawa (Data Analysis Fusion WG)
Computational Science Research Program, RIKEN Hisayuki Miyashita (Molecular Scale WG) - Winter School 2011 for the Integrated Simulation of Living Matter
