
Brain and Neural WG
Whole Brain Simulation
of the Insect Olfactory System

Research Center for Advanced Science and Technology,
The University of Tokyo
Tomoki Kazawa, Stephan Shuichi Haupt
Nerve cells (neurons) are the building blocks necessary for the emergence of intelligence as they process internal and external information, generally encoded in sequences of spikes and form networks in which such information is relayed between them. Input and output parameters of neurons such as gain can change dynamically depending on prior activity history. Neurons are present in all metazoan animals from simple coelenterates like Hydra and have the same general properties across the animal kingdom. Though the basic building blocks are so similar, there is a big difference in the size of nervous systems, human brains harbour 1011-12 neurons, insects have 105-6 brain neurons, and the somatic nervous system of the nematode Caenorhabditis elegans contains just 302 neurons. Insects are sufficiently complex to display many sophisticated behaviours, such as learning floral odors and colors. Multimodal information is integrated in the insect brain which can generate a wide variety of complex and plastic behaviours, making insect brains a highly suitable system to investigate the emergence of intelligence. Our approach to understanding such an intelligent system shaped by millions of years of evolution is to attempt to recreate it through simulation. This insect whole brain simulation has to encompass information processing from sensory input to behavioural ouput. We currently focus on the odour-source localisation behaviour of silkmoth (Bombyx mori) males as a convenient model system. We are analyzing the brain of the silkmoth accumulating numerous data obtainted with a large grid of methods including molecular genetics, electrophysiology, imaging, immunohistochemistry, and behavioural experiments. Our methods cover the full scale of observation levels from the molecular and cellular levels to the network and behaviour levels. These multi-scale data are integrated in our database, which, for example, contains data on >1600 neurons individually identified in terms of 3D-morphology and physiology, currently making it the largest single-species individual neuron database worldwide. An important insight from these data was the understanding of global and regional connectivity in the olfactory system in the brain of the silkmoth.
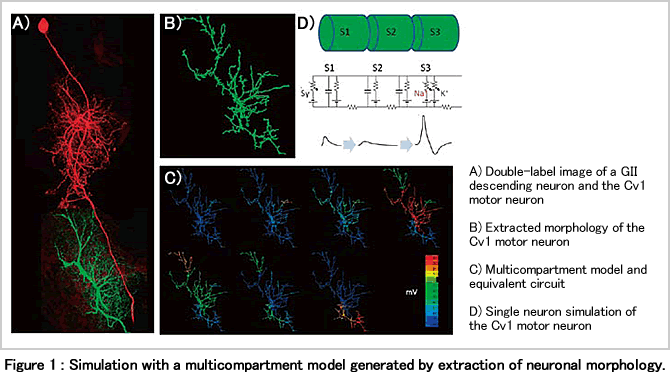
Since insects can exhibit highly complex behaviours despite having a relatively small number of neurons, it is likely that the relative contribution of each individual neuron is more important than for instance in mammals. In our largescale simulation of insect neural circuits, we are therefore performing a detailed simulation of each individual neuron using multicompartment models (Figure 1). The 3D-morphology of neurons is acquired by confocal microscopy and can then be approximated by sets of connected cylinders. A cylinder per se has a simple equivalent circuit consisting of voltage and impedance to the outside of the cylinder (membrane resistance and capacitance). Cylinders are connected by resistors that correspond to axial resistance in neurites. Inputs are described as chemical synapses having postsynaptic potentials of 5-20ms duration upon the simulated activation by neurotransmitter. The synaptic activation can induce spikes of 1-2ms width that are modeled by the conventional sodium and potassium currents (Figure 1C). By mapping and combining numerous single neuron simulation models in the brain, a neural circuit simulation can be constructed (Figure 2).
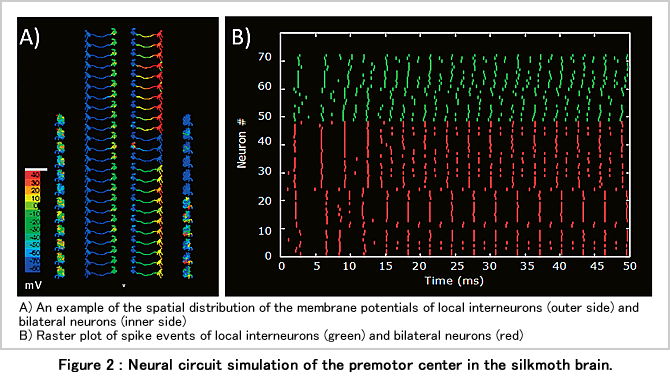
We estimate that a total of about 10000 neurons in the brain are involved the whole information processing from reception of olfactory sensory input to the generation of the control activity patterns for odour-source localisation behaviour. Using representations of neurons consisting of about 100 compartments, we estimated that the computations necessary to simulate information processing in the entire insect brain can be performed in real time on a machine with the performance of a petaflop HPC. If a real-time simulation of the neural circuit from sensory input to motor output can be achieved, it also becomes possible to investigate information processing by modification of the circuit properties in ways that are currently not feasible in actual experiments. Such manipulations are of paramount importance to identify minimal neural circuits to perform specific information processing tasks. We expect this approach will have an impact in medical applications, in particular neurorehabilitation and neural protheses.
BioSupercomputing Newsletter Vol.3
- SPECIAL INTERVIEW
- The role of supercomputers is important for integrating various leading-edge research bases for proactive use.
Manager Clinical Development Planning and Management
Mochida Pharmaceutical Co., Ltd. Visiting Professor Tohoku University Kazumi Nishijima - Sonic simulation research in the body which is essential for promotion of ultrasound therapy and development
of therapeutic apparatus
Extraordinary researcher, Department of Mechanical Engineering, School of Engineering, University of Tokyo Akira Sasaki
- Report on Research
- Achievement of a Multiscale Molecule Simulation of QM/MD/CGM(Molecular Scale WG)
Institute for Protein Research, Osaka University
Yasushige Yonezawa / Shusuke Yamanaka / Hiromitsu Shimoyama / Hideki Yamazaki / Haruki Nakamura
RIKEN, Computational Science Research Program Ikuo Fukuda - Towards development and experimental demonstration of liver model based
on large-scale metabolic simulation at individual cellular level (Cell Scale WG)
School of Medicine, Keio University Ayako Yachie-Kinoshita - Exhaustive Protein-Protein Interaction Network Prediction by Using MEGADOCK (Data Analysis Fusion WG)
Graduate School of Information Science and Engineering, Tokyo Institute of Technology
Yutaka Akiyama / Yuri Matsuzaki / Nobuyuki Uchikoga / Masahito Ohue - Whole Brain Simulation of the Insect Olfactory System
Research Center for Advanced Science and Technology, The University of Tokyo Tomoki Kazawa, Stephan Shuichi Haupt
- Report
- The summer school 2010 for the Integrated Simulation of Living Matter was held.
RIKEN, Computational Science Research Program Yasuhiro Ishimine (Organ and Body Scale WG)
The Institute of Medical Science, The University of Tokyo Teppei Shimamura (Data Analysis WG)
RIKEN, Computational Science Research Program Yasuhiro Sunaga (Cell Scale WG)
Kyoto University Graduate School of Informatics Naoki Honda (Brain and Neural WG)
RIKEN, Computational Science Research Program Gen Masumoto (High-Performance Computing Team)
RIKEN, Computational Science Research Program Kosuke Matsunaga (Molecular Scale WG) - After participation in the summer school 2010 for the Integrated Simulation of Living Matter
First year of doctor's course, The University of Tokyo Graduate School of Science Ken Saito
