
Cell Scale WG
Towards development and experimental
demonstration of liver model based on large-scale metabolic simulation
at individual cellular level

School of Medicine, Keio University
Ayako Yachie-Kinoshita
Hepatic parenchymal cells (hepatocytes) which occupy a large part of the cellular volume in liver have very extensive and complex metabolic pathways including most metabolic reactions occurring in the body, and play an important role in regulation of vast variety of small molecules. Compartmentation of metabolism by site specificity, in which an enzyme expression pattern is different between the upstream (portal vein) and downstream (central vein) of sinosidal blood flow, is considered to be one of the factors that makes the functional switching of metabolism in liver efficient. However, due to the striking complexity in metabolic dynamics in which many metabolic functions are intricately involved with each other, it is very difficult to comprehensively understand the dynamics of hepatic metabolism and its underlying mechanism, and for many phenomena, analogies are still being drawn from circumstantial evidence. In such a case, model-based analysis and simulation by integration of independent kinetic characteristics is very powerful to understand the system as a whole.
We therefore established an unprecedented large-scale metabolic model including approximately 500 defined substances and about 250 dynamic reactions using the E-Cell Simulation EnvironmentNote1 based on the reaction kinetics obtained by literature search. With this model, the dynamics of the metabolism in the mitochondria and cytoplasm can be predicted separately and simultaneously(considering the compartmentation of metabolism in the cells), and the effects of exchange of small molecules between hepatocytes in portal vain and central vein regions via a gap junction can be predicted by changing the enzyme expression pattern to set up the regional heterogeneity of the both regions (considering the compartmentation of metabolism between cells). We are also developing the model to reproduce the condition in which hepatocytes with different metabolic properties interlock each other to function by using the Riken Cell Simulator (RICS)Note2 (Figure 1).
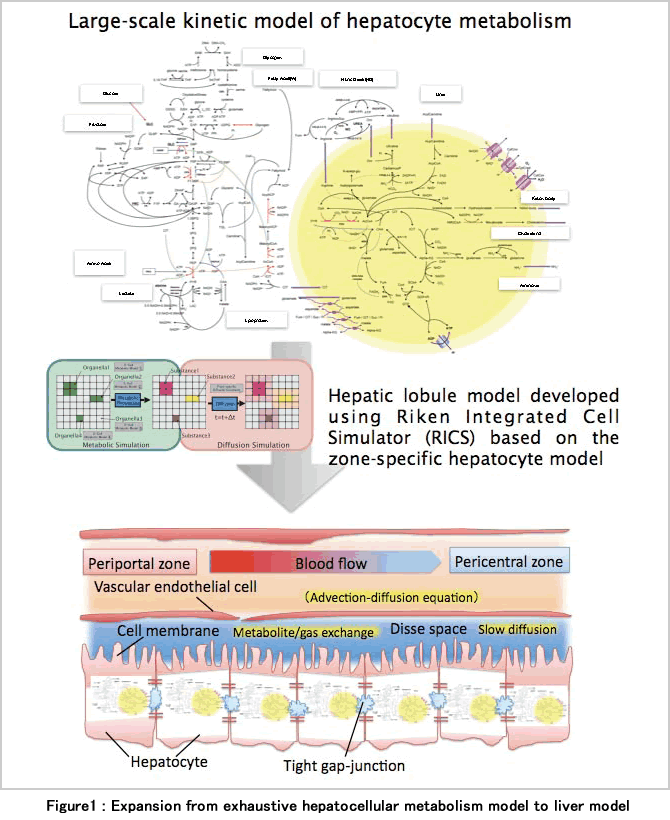
Using the large-scale metabolic model of hepatocyte, we compared a predicted dynamics under hypoxic and hypoglycemic condition from a couplet model in which two connecting hepatocytes having portal vein and central vein metabolic properties with those from the models in which both hepatocytes have a property of portal vein or central vein only. As a result, by coupling cells with different properties, over-all changes in metabolites and a decrease in ATP energy in response to these stimulations were suppressed, and it was predicted that the robustness of the metabolism would be improved at an organ level.
This result may contribute not only to understanding how hepatic metabolism is optimized using heterogeneity, but also to understanding of metabolic failure in the intercellular material exchange disorder such as a case of hepatic cirrhosis. Moreover, the in vivo experiments in our laboratory have clearly showed that the metabolism changed largely not only in metastatic foci but also the surrounding hepatocytes due to hepatic metastasis of colon cancer. Our hepatic metabolism simulation model can be applied to reproduction and discussion of such a disease model, and prediction of resistance to hepatic ischemia during surgery and spare ability at hepatectomy.
In considering the effects of cell metabolism on organs and the whole body, particularly pathological conditions, on the other hand, it may be essential to establish an experimental system to observe the metabolic conditions without destroying the tissue structure.
Our laboratory therefore focus on an experimental technique of “visualization of metabolism with μm resolution” combining a metabolome analysis with an imaging mass spectrometryNote3 (Hattori K et al., Antioxid Redox Signal. 13(8): 1157-67 (2010)). At present, improvement of this method is moving ahead with great urgency, and it has become possible to determine more metabolites including aminoacids at higher resolution. By this technique, it may be possible to measure and quantify the difference in metabolism between portal vein and central vein hepatocytes with the in vivo spatial arrangement (Figure 2).
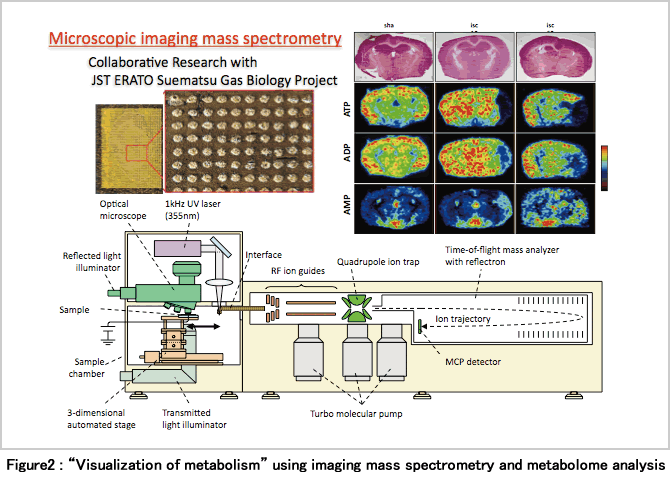
In the future, the study to quantify and observe metabolism as the in vivo spatial arrangement of tissue is maintained, and to elucidate the results and the meaning of material variation by exhaustive cell simulation (large-scale calculation with a supercomputer) considering spatial information will become important.
Note 1: Simulation environment focuses on phenomena of intracellular molecules, which has been being developed by Institute for Advanced Biosciences of Keio University, and RIKEN.
Note 2: A platform of cell simulation, which is being developed by RIKEN.
The reactions of small molecules considering the spatial information in the cell and the position among cells can be simulated by constructing the cellular model in the space (voxel) of a fixed grid, and establishing diffusion and transfer among voxels.
Note 3: The mass distribution is visualized at a resolution of 10 μm by ionizing metabolites by applying a laser to a section of frozen tissue at fine intervals, by measuring ions with a mass spectrometer connected to it, and then by reconstructing information on masses of the metabolites determined at the respective positions.
BioSupercomputing Newsletter Vol.3
- SPECIAL INTERVIEW
- The role of supercomputers is important for integrating various leading-edge research bases for proactive use.
Manager Clinical Development Planning and Management
Mochida Pharmaceutical Co., Ltd. Visiting Professor Tohoku University Kazumi Nishijima - Sonic simulation research in the body which is essential for promotion of ultrasound therapy and development
of therapeutic apparatus
Extraordinary researcher, Department of Mechanical Engineering, School of Engineering, University of Tokyo Akira Sasaki
- Report on Research
- Achievement of a Multiscale Molecule Simulation of QM/MD/CGM(Molecular Scale WG)
Institute for Protein Research, Osaka University
Yasushige Yonezawa / Shusuke Yamanaka / Hiromitsu Shimoyama / Hideki Yamazaki / Haruki Nakamura
RIKEN, Computational Science Research Program Ikuo Fukuda - Towards development and experimental demonstration of liver model based
on large-scale metabolic simulation at individual cellular level (Cell Scale WG)
School of Medicine, Keio University Ayako Yachie-Kinoshita - Exhaustive Protein-Protein Interaction Network Prediction by Using MEGADOCK (Data Analysis Fusion WG)
Graduate School of Information Science and Engineering, Tokyo Institute of Technology
Yutaka Akiyama / Yuri Matsuzaki / Nobuyuki Uchikoga / Masahito Ohue - Whole Brain Simulation of the Insect Olfactory System
Research Center for Advanced Science and Technology, The University of Tokyo Tomoki Kazawa, Stephan Shuichi Haupt
- Report
- The summer school 2010 for the Integrated Simulation of Living Matter was held.
RIKEN, Computational Science Research Program Yasuhiro Ishimine (Organ and Body Scale WG)
The Institute of Medical Science, The University of Tokyo Teppei Shimamura (Data Analysis WG)
RIKEN, Computational Science Research Program Yasuhiro Sunaga (Cell Scale WG)
Kyoto University Graduate School of Informatics Naoki Honda (Brain and Neural WG)
RIKEN, Computational Science Research Program Gen Masumoto (High-Performance Computing Team)
RIKEN, Computational Science Research Program Kosuke Matsunaga (Molecular Scale WG) - After participation in the summer school 2010 for the Integrated Simulation of Living Matter
First year of doctor's course, The University of Tokyo Graduate School of Science Ken Saito
