
Protein Reaction Simulation Based
on All-electron Calculation
Institute of Industrial Science, the University of Tokyo (Molecular Scale WG)
(From left)Fumitoshi SATO, Toshiyuki HIRANO, Noriko UEMURA, Naoki TSUNEKAWA, Junichi MATSUDA

Our group belongs to the Molecular Scale Team, which is trying to obtain a better understanding of the molecular world – the foundation of the activities of life – and is in charge of the most fundamental measures. For details of the team, please see the article written by Akinori Kidera, the team leader, in the previous issue.
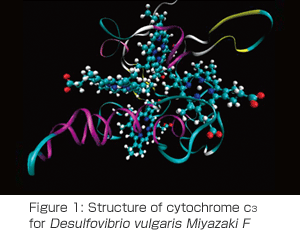
The goal of our research is, in a word, to develop allelectron simulation software for proteins, which will bring out the best performance in the next-generation supercomputer. As target software, we selected ProteinDF. This software has achieved all-electron wave function calculation for various proteins using the densit y functional theory that is a standard electronic state calculation method. Toshiyuki Hirano is in charge of the migration of ProteinDF to ISLiM and the tuning of massive parallelization. He plays a key role in expanding the ability and functions of ProteinDF and will develop amazing simulations that make the best use of the next-generation supercomputer. Junichi Matsuda is a professional that has tuned various programs on various processors and joined our team on November 1, 2009. His assignment was about to be excluded from the budget for the next fiscal year two weeks after first participating, but he was able to join us as planed to offer his best in tuning the programs for the Venus chip. For the electronic state calculation, we need a technology that succeeds self-consistent calculation under complicated conditions, as well as brings out hardware performance. The required task for this is sort of like controlling the space shuttle from launch to landing and the larger the molecule is, the more difficult it becomes. The person in charge of this task is a specialist, Noriko Uemura. As a touchstone of her newly developed method, she is grappling with the calculation of the cytochrome C3 molecule, which has four hemes (iron-porphyrin) in a single body and is the motif for the ProteinDF logo. (Figure 1). Finally, Naoki Tsunekawa is the researcher that is taking on the challenge of the free-energy calculation using allelectron calculation. As a strategy, he is trying to connect different energy functionals. To prove the validity of this strategy, he would not mind tackling several million quantum chemical calculations (and he actually has).
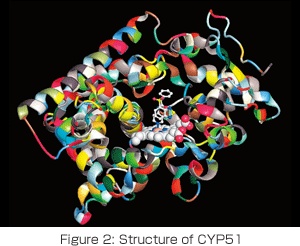
Having these members, we have been developing software that helps us understand protein molecules from an electronic state by bringing out the best in the nextgeneration supercomputer. We will put our energy into not only basic research, but also into research for more practical applications, such as a drag-resistance simulation for cytochrome P450 (CYP) (Figure 2). We also have great expectations for applications to energy and environmental issues, such as the CO2 fixation of RubisCo, artificial photosynthesis and biomimetic industrial catalysts. Please join us in our research.
ProteinDF_ISLiM has already succeeded in obtaining an MPI/OpenMP hybrid parallel calculation structure and achieved the calculation in 2,500 parallel cores using a Cray XT-5. As a result of this achievement, ProteinDF_ISLiM was chosen as one of the first runner applications. We are currently in the development phase where ProteinDF_ ISLiM is being optimized for the open specifications of the next-generation supercomputer. Helped by the knowledge from RIKEN and Fujitsu Ltd., we are going to catch up with the hardware development process. Please look forward to our future activities.
BioSupercomputing Newsletter Vol.2
- SPECIAL INTERVIEW
- Aiming to Become a Global Trendsetter in the Life Sciences by Making the Best Use of the Next-generation Supercomputer!
Computational Science Research Program Deputy Program Director
Ryutaro HIMENO
- A Message from the Team Leader
- Simulating Cell Phenomena by Recreating Cells as They Exist in Living Organisms
Cell Scale Team Team Leader
Hideo YOKOTA - Create a Brain on a Supercomputer to Unravel the Functions of the Brain and Nervous System
Brain and Neural Systems Team Team Leader
Shin ISHII - High-performance Computing Environment to Maximize the Potential of the Next-generation Supercomputer
High-performance Computing Team Team Leader
Makoto TAIJI
- Report on Research
- Protein Reaction Simulation Based on All-electron Calculation
Institute of Industrial Science, the University of Tokyo
Fumitoshi SATO / Toshiyuki HIRANO / Noriko UEMURA / Naoki TSUNEKAWA / Junichi MATSUDA - Full Eulerian Fluid-structure Coupled Method
Associate Professor at School of Engineering, the University of Tokyo
Kazuyasu SUGIYAMA - A Large-scale Simulation Model of Cortical Microcircuits: CMDN(Cortical Microcircuit Developed on NEST)
Brain and Neural System Team
Jun IGARASHI - Development of Next-generation Molecular Dynamics Simulation Programs
High-performance Computing Team
Hiroshi KOYAMA / Yosuke OHNO / Gen MASUMOTO / Aki HASEGAWA / Gentaro MORIMOTO
