
Cell Scale Team
Simulating Cell Phenomena by
Recreating Cells as They Exist in
Living Organisms

Cell Scale Team
Team Leader
Hideo YOKOTA
There are many approaches that can be applied to theoretical biology, ranging from the micro-level method based on molecular dynamics calculation to macro-level methods such as mechanical simulation. Our cell research takes an approach that falls under the latter category. Our ongoing research in the digitization of a living organism's shape information is now at the stage where whole-body profile data of a mouse has been obtained at a 10-µm resolution. Furthermore, advances in live-cell imaging technology have enabled us to collect and digitize the data of a living cell and study its dynamic process. This means a variety of cell phenomena, including cell shapes, can be now observed as four-dimensional digitized data. Imaging technology, however, is not without its limits. Accordingly, many intracellular phenomena remain unexplainable. Our team is working on the development of virtual cell modeling, which will allow us to understand what happens inside a cell through digital simulation.
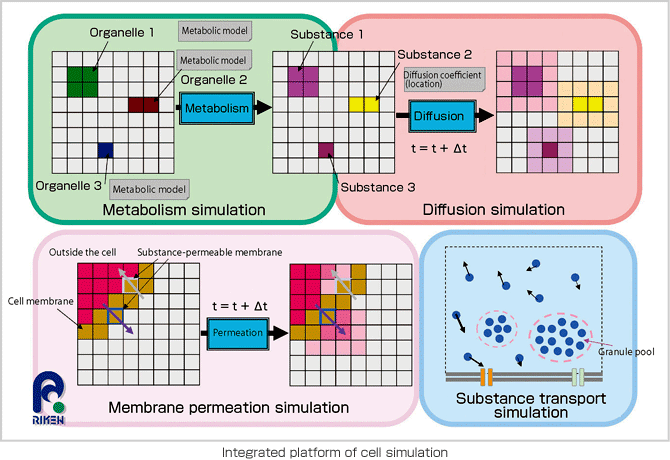
What are the key factors in understanding cell phenomena? The answers are intracellular chemical reaction networks and membranes that serve as the structural components of the cell. The computing of chemical reactions (i.e., metabolism) incurs enormously-high costs, although in the future molecular dynamics-based calculation may be applied. To bypass this problem, we intend to create a simulated cell model by using certain chemical reactions that appear to be dominant in the cell. There are several precedent studies in this field, such as the E-CELL system by the Institute for Advanced Biosciences at Keio University and the Cell Illustrator by the Human Genome Center at the University of Tokyo. However, these systems simulate chemical reactions by assuming the cell to be a closed bag filled with homogeneous materials or by making a cell model that is extremely simplified, thus ignoring the cell's location. Living cells have shapes and organelles, which perform different vital functions. The reactions of organelles, such as Golgi bodies and mitochondria, vary depending on the environment. As such, simulation requires an environmental context, which will allow us to recreate cells on a computer as they exist in a living orgasm. In doing so, we also need to be able to describe the diffusion of substances and the cytoskeletonbased active transport process in addition to intracellular chemical reactions. The functions of cell membranes, the other key factor, include a channel that allows specific ions to permeate, a pump that moves substances by expending energy called ATP and a receptor that receives signals from outside the cell. Our aim is to create a comprehensive simulation model that addresses all factors, including the cell's location, in an effort to understand all the phenomena that occur within living cells.
Another important note on the cell’s location is the fact that a cell does not exist as an independent unit. Cells in the liver, for example, each have roles to play according to their location, enabling the liver to perform its functions properly. The location within the organ also determines types of cell reactions among the liver cells. Consequently, a simulation that ignores this factor lacks merit when attempting to replicate living cells influenced by their surrounding environment.
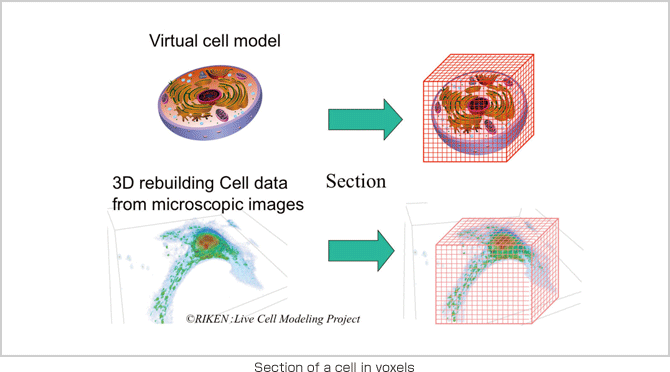
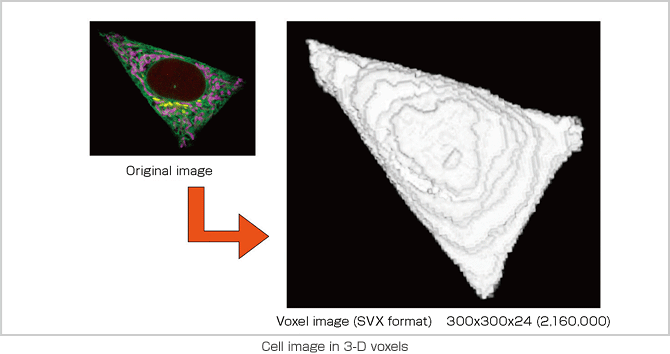
In order to conduct a simulation with a cell model that authentically replicates cell behavior in a living organism, we are working on the development of application software called RICS. The software creates a cell model in a lattice space with 100-nm sections and applies experiment data on intracellular phenomena to conduct coupled calculation of chemical reactions, substance diffusion and membrane permeation on a one millionvoxel volumetric display (1003). The system is designed to be a common simulation platform, inclusive of all types of cells. A development project currently targeting hepatocytes is planned to shift its focus onto hepatic lobule simulation next. However, simulation itself is not the goal; rather, it is a tool that will lead us to an understanding of the phenomena and functions of life. It is also important that these simulations contribute to medical science. We aim to carry our development further to the simulation and analysis of cell reactions to drugs and the removal of organs and cancerous tumors. Potential applications of the simulation model also include the design of artificial and regenerated organs, for which the calculation of cell locations is critical.
BioSupercomputing Newsletter Vol.2
- SPECIAL INTERVIEW
- Aiming to Become a Global Trendsetter in the Life Sciences by Making the Best Use of the Next-generation Supercomputer!
Computational Science Research Program Deputy Program Director
Ryutaro HIMENO
- A Message from the Team Leader
- Simulating Cell Phenomena by Recreating Cells as They Exist in Living Organisms
Cell Scale Team Team Leader
Hideo YOKOTA - Create a Brain on a Supercomputer to Unravel the Functions of the Brain and Nervous System
Brain and Neural Systems Team Team Leader
Shin ISHII - High-performance Computing Environment to Maximize the Potential of the Next-generation Supercomputer
High-performance Computing Team Team Leader
Makoto TAIJI
- Report on Research
- Protein Reaction Simulation Based on All-electron Calculation
Institute of Industrial Science, the University of Tokyo
Fumitoshi SATO / Toshiyuki HIRANO / Noriko UEMURA / Naoki TSUNEKAWA / Junichi MATSUDA - Full Eulerian Fluid-structure Coupled Method
Associate Professor at School of Engineering, the University of Tokyo
Kazuyasu SUGIYAMA - A Large-scale Simulation Model of Cortical Microcircuits: CMDN(Cortical Microcircuit Developed on NEST)
Brain and Neural System Team
Jun IGARASHI - Development of Next-generation Molecular Dynamics Simulation Programs
High-performance Computing Team
Hiroshi KOYAMA / Yosuke OHNO / Gen MASUMOTO / Aki HASEGAWA / Gentaro MORIMOTO
